- Blog
- Music math games for kids
- Adobe flash cs3 pixel tools
- Im pro golfer drug commercial
- Download buildbox
- Timing chain symptoms
- Easycatalog pricing
- Iphone xs max starcraft ii images
- Hospice nurse reveals unexplained phenomena
- Daylife projects
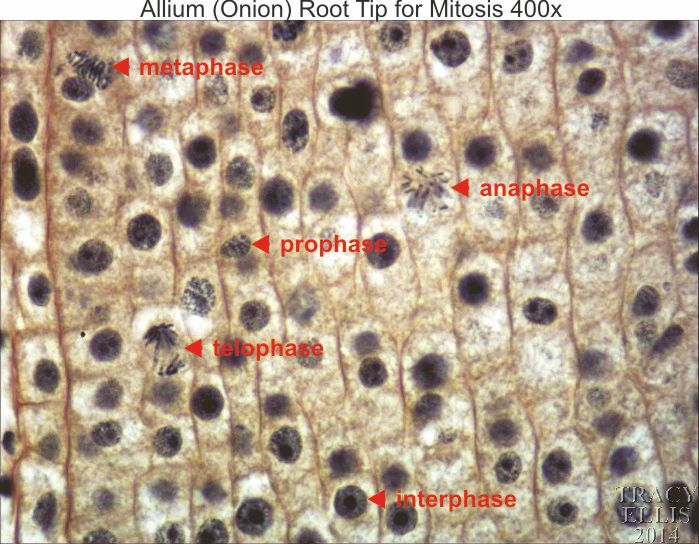
- Prophase under microscope
- Capto core 400
- Cinesync and rv
- Traktor vs serato
_Pressed;_root_meristem_of_Vicia_faba_(cells_in_anaphase,_telophase).jpg)
Subsequent DNA repair results in either crossover (CO) or non-crossover (NCO) events (reviewed in Osman et al., 2011 Hunter, 2015 Mercier et al., 2015 Sansam and Pezza, 2015), according to the selected repair template and the resolution of repair intermediates. The break ends are resected and coated with the RecA-related recombinases RAD51 and DMC1, highly conserved proteins that facilitate strand invasion of homologous sequences ( Bishop, 1994 Dresser et al., 1997 Doutriaux et al., 1998 Couteau et al., 1999 Li et al., 2004 Kurzbauer et al., 2012). Once the DSB has been formed, SPO11 is released from the DNA by the MRE11/RAD50/Xrs2-NBS1 (MRX/N) complex stimulated by COM1/Sae2 ( Neale et al., 2005 Uanschou et al., 2007 Milman et al., 2009 Cannavo et al., 2018). The meiotic axis is formed by axial element proteins like ASY1/Hop1, ASY3/Red1, and ASY4 ( Hollingsworth et al., 1990 Rockmill and Roeder, 1990 Armstrong et al., 2002 Ferdous et al., 2012 Chambon et al., 2018 West et al., 2019), together with cohesin proteins, among them SCC3 and REC8 ( Klein et al., 1999 Toth et al., 1999 Cai et al., 2003 Chelysheva et al., 2005), and it is required for several processes from DSB formation to recombinational repair. SPO11-interacting proteins link sites of DSB-formation to the chromosome axis ( Blat et al., 2002 Panizza et al., 2011 Acquaviva et al., 2013). Several proteins necessary for DSB formation have been identified in a variety of organisms, with the conserved topoisomerase-related protein SPO11 as the catalytically active factor within the DSB-forming complexes (reviewed in Keeney, 2001 Edlinger and Schlogelhofer, 2011 Lam and Keeney, 2014 Robert et al., 2016). The coordinated and tightly controlled formation of DNA double-strand breaks (DSBs) and their repair is a prerequisite for successful meiotic divisions: it ensures the pairing and segregation of homologous chromosomes as well as re-shuffling of genetic traits. Meiosis is completed by the formation of four genetically different haploid precursor cells that develop into gametic cells. Thereafter, sister chromatids segregate during the second division. During the first meiotic division, after DNA replication, homologous chromosomes pair, recombine and are then separated to opposite poles of the cell. In contrast to somatic cells that give rise to identical daughter cells by mitotic (equational) cell division, germ cells divide meiotically to form haploid gametes, thereby ensuring constant karyotypes over generations. Understanding its molecular mechanisms and involved factors is therefore essential for human health and fertility and, importantly, for plant breeding and food security. Meiosis is a specialized cell division and the basis for genetic diversity through sexual reproduction. We review different techniques, focusing on stimulated emission depletion (STED) nanoscopy, to offer researchers guidance for selecting the optimal protocol and equipment to address their scientific question. Here, we provide an overview of classical and advanced sample preparation and microscopy techniques with an updated Arabidopsis meiotic atlas based on super-resolution microscopy.
New technologies enable observation of cells and nuclei at a nanometer scale and hold great promise to the field since they allow observing complex meiotic molecular processes with unprecedented detail. Recent advances in super-resolution technologies changed how microscopic images are acquired and analyzed. Visualization of meiotic chromosomes and the proteins involved in meiotic recombination have become essential to study meiosis in many systems including the model plant Arabidopsis thaliana. Department of Chromosome Biology, Max Perutz Labs, University of Vienna, Vienna BioCenter, Vienna, Austria.Jason Sims * †, Peter Schlögelhofer † and Marie-Therese Kurzbauer * †
